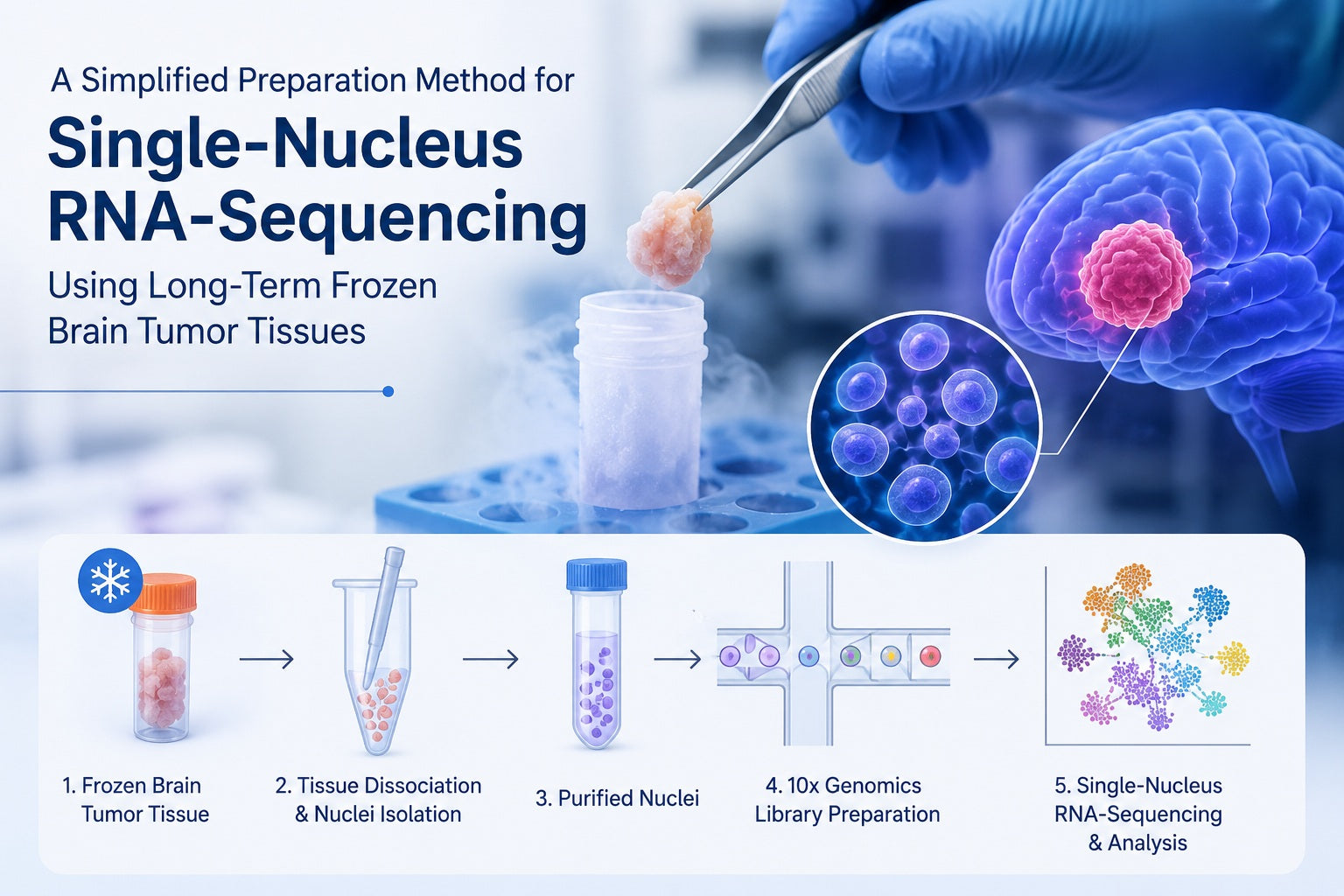
Single-nucleus RNA-sequencing is a sequencing approach that profiles gene expression from isolated nuclei instead of intact whole cells. It is especially useful for frozen, archived, fragile, or hard-to-dissociate tissues, including brain tumor tissue. For researchers studying pediatric glioma, tumor heterogeneity, and complex tumor microenvironments, a simplified preparation method can help transform long-term frozen specimens into usable nuclear suspensions for snRNA-seq, single-cell sequencing analysis, and downstream molecular biology research.
How Can Long-Term Frozen Brain Tumor Tissue Be Prepared for snRNA-seq?
Long-term frozen brain tumor tissue can be prepared for snRNA-seq by keeping samples cold, gently disrupting the tissue, lysing cell membranes while preserving nuclei, filtering and washing the nuclear suspension, removing debris when needed, counting intact nuclei, and preparing nuclei at the correct concentration for sequencing. A successful nucleus isolation protocol focuses on nuclear integrity, low debris, consistent recovery, and compatibility with downstream NGS library preparation.
What Is Single-Nucleus RNA-Sequencing?
Single-nucleus RNA-sequencing, often called snRNA-seq, measures RNA transcripts from individual nuclei. Unlike single-cell RNA-sequencing, which requires viable intact cells, snRNA-seq can work with frozen tissue where whole-cell isolation may be challenging. This makes it valuable for archived research samples, biobanked tumor specimens, neural tissues, and dense or delicate samples that may not produce high-quality single-cell suspensions.
In brain tumor research, snRNA-seq helps researchers study cellular diversity across malignant, immune, stromal, vascular, and neural-associated populations. It can support deeper exploration of tumor heterogeneity, lineage states, microenvironment interactions, and gene expression patterns in pediatric glioma and other brain tumor models.
Why Does snRNA-seq Matter for Frozen Brain Tumor Workflows?
Frozen brain tumor tissues are important research resources. Many high-value samples are stored long-term because fresh tissue processing is not always practical at the time of collection. snRNA-seq expands the usefulness of these samples by allowing researchers to isolate nuclei from frozen material rather than relying only on fresh single-cell RNA-sequencing workflows.
For modern labs, this matters in several ways:
- It supports research using archived and biobanked brain tumor tissue
- It helps profile fragile tissues that are difficult to dissociate into intact cells
- It provides access to nuclear transcriptomes from rare tumor samples
- It supports studies of pediatric glioma and tumor heterogeneity
- It can align with downstream NGS library preparation and computational analysis
- It helps teams build more flexible single-cell sequencing strategies
A practical workflow should protect nuclear integrity while producing a clean, countable suspension suitable for sequencing platform input.
Key Applications of snRNA-seq Using Frozen Brain Tumor Tissues
Pediatric Glioma Research
Pediatric glioma samples can be limited, precious, and difficult to process as fresh tissue. A nuclear isolation protocol for pediatric glioma gives researchers a way to work with frozen specimens while preserving cell-type information at single-nucleus resolution.
Tumor Heterogeneity Studies
Brain tumors contain multiple transcriptional states and diverse cell populations. snRNA-seq helps researchers explore malignant cell states, immune infiltration, glial-associated populations, endothelial cells, stromal signals, and sample-to-sample variability.
Single-Cell Sequencing of Frozen Tumor Samples
When whole-cell dissociation is difficult, snRNA-seq provides a practical route for single-cell sequencing of frozen tumor samples. This is useful for dense, necrotic, fragile, or long-term stored tissues.
NGS Library Preparation and Molecular Profiling
Once high-quality nuclei are isolated, downstream library preparation enables transcriptomic profiling. Nucleus quality, concentration, debris level, and RNA preservation all influence downstream assay performance.
Translational and Diagnostic Research Workflows
In research settings, frozen tissue compatibility can help laboratories study valuable sample collections. While snRNA-seq is not a standalone clinical diagnostic claim, it supports molecular research, biomarker exploration, and tumor biology studies.
Simplified Workflow: Preparing Frozen Brain Tumor Tissue for snRNA-seq
A simplified workflow should be consistent, cold, gentle, and compatible with downstream sequencing. Exact conditions vary by tissue type, tumor condition, storage history, and platform requirements, so teams should follow validated protocols and kit instructions.
Step 1: Review Sample Quality and Storage History
Start by documenting the tissue source, storage duration, freezing method, approximate tissue mass, tumor region, and any known necrotic or hemorrhagic areas. Long-term frozen brain tumor tissue can vary in texture, cellularity, and debris content. Reviewing sample history helps the team choose a realistic workflow and set appropriate recovery expectations.
Step 2: Keep Tissue and Reagents Cold
Cold handling supports nuclear integrity and RNA preservation. Prepare buffers, tubes, strainers, centrifuge settings, and cleanup reagents before removing samples from storage. A well-organized cold workflow reduces handling time and helps operators move smoothly through each step.
Step 3: Gently Disrupt the Frozen Tissue
Frozen brain tumor tissue may require careful mechanical disruption before lysis. The goal is to expose nuclei without excessive shear. Avoid over-processing. Gentle chopping, douncing, or kit-directed disruption can help release nuclear material while preserving nuclear morphology.
Step 4: Lyse Cell Membranes While Preserving Nuclei
The core of a nucleus isolation protocol is selective lysis. The buffer should disrupt cell membranes while keeping nuclei intact. Brain and tumor samples may contain lipid-rich material, extracellular matrix, dead cell debris, and variable cellular density, so lysis conditions should be optimized for the sample type.
Step 5: Filter and Wash the Nuclear Suspension
Filtration removes tissue fragments and large debris. Washing helps reduce background material and prepare nuclei for counting or cleanup. The number of washes should balance cleanliness and recovery. Too few washes may leave debris, while excessive handling can reduce yield. A simplified method should include clear checkpoints for visual inspection and counting.
Step 6: Remove Debris When Needed
Frozen tumor samples may contain debris from necrotic regions, lipid-rich brain components, and damaged cells. Debris removal or nuclear cleanup can improve suspension quality for sequencing input. For labs working with pediatric glioma or long-term frozen tumor tissue, cleanup steps can be especially helpful when the starting material is limited or variable.
Step 7: Count Nuclei and Assess Quality
Before library preparation, count nuclei and assess suspension quality. Useful checkpoints include:
- Intact nuclei appearance under microscopy
- Nuclei concentration
- Debris level
- Clumping level
- Recovery from the starting material
- Compatibility with sequencing platform input requirements
A clean, countable suspension supports better workflow efficiency and more confident downstream analysis.
Step 8: Prepare for NGS Library Preparation
Once nuclei are prepared at the correct concentration, proceed according to the chosen snRNA-seq platform and NGS library preparation workflow. Keep documentation complete, including sample ID, tissue mass, buffer lot, incubation timing, wash count, operator, and final nuclei concentration.
snRNA-seq vs Single-Cell RNA-Sequencing for Frozen Tumor Samples
Both single-cell RNA-sequencing and snRNA-seq are powerful, but they fit different sample realities.
|
Workflow |
Best Fit |
Key Consideration |
|
Single-cell RNA-sequencing |
Fresh tissue with recoverable intact cells |
Requires viable cells and optimized tissue dissociation |
|
Single-nucleus RNA-sequencing |
Frozen, archived, fragile, or hard-to-dissociate tissue |
Requires intact nuclei, low debris, and suitable nuclear cleanup |
|
Bulk RNA-seq |
Average gene expression from mixed tissue |
Does not resolve individual cellular populations |
|
Spatial transcriptomics |
Preserves tissue location context |
Often complements, rather than replaces, single-nucleus profiling |
For long-term frozen brain tumor tissues, snRNA-seq often provides a practical path when fresh tissue dissociation is not available.
Important Factors to Consider Before Buying
|
Buying Factor |
Why It Matters |
Suggested Category |
|
Frozen tissue compatibility |
Long-term stored samples require gentle nuclear release |
|
|
Brain or tumor tissue type |
Brain tumors may include lipid-rich, necrotic, and heterogeneous regions |
Neuroscience Single-Cell Sample Prep / Tumor Sample Prep |
|
Debris removal |
Cleaner suspensions support counting and sequencing input |
Sample Preparation Kits |
|
Whole-cell comparison |
Some studies compare scRNA-seq and snRNA-seq results |
Tissue Dissociation Kits |
|
Downstream sequencing |
Nuclei quality supports library preparation and analysis |
NGS Library Prep Kits |
|
Molecular follow-up |
Extracted material may support PCR, qPCR, or other assays |
PCR/qPCR and Molecular Biology Reagents |
|
QC workflow needs |
Documentation and reagent consistency support reproducibility |
General Research Reagents and QC Tools |
Practical Tips for Better Results
A strong snRNA-seq preparation method is built on planning, consistency, and quality checks.
Helpful workflow practices include:
- Prepare buffers, tubes, strainers, and equipment before sample retrieval
- Keep tissue and reagents cold throughout the workflow
- Use gentle disruption to preserve intact nuclei
- Avoid unnecessary handling after nuclei are released
- Filter suspensions to remove large debris
- Include cleanup steps when debris is visible, or counting is difficult
- Count nuclei carefully before sequencing input
- Document tissue mass, storage history, reagent lots, wash steps, and final concentration
- Run pilot preparations when working with a new brain tumor type or storage condition
These practices support more consistent snRNA-seq using frozen brain tumor tissues and help teams compare results across samples.
FAQs
What is single-nucleus RNA-sequencing?
Single-nucleus RNA-sequencing is a method that profiles RNA from individual nuclei rather than intact cells. It is useful for frozen, archived, fragile, or hard-to-dissociate tissues where whole-cell recovery may be challenging.
Can snRNA-seq be used with long-term frozen brain tumor tissue?
Yes. snRNA-seq is well-suited for many frozen tissue workflows because nuclei can often be isolated from samples that are not ideal for whole-cell dissociation. The workflow should focus on cold handling, intact nuclei recovery, debris reduction, and accurate counting.
Why is snRNA-seq useful for pediatric glioma research?
Pediatric glioma samples can be rare and valuable. snRNA-seq helps researchers study cellular diversity and tumor heterogeneity from frozen tissue, supporting research into gene expression patterns and tumor microenvironment composition.
What is the difference between scRNA-seq and snRNA-seq?
Single-cell RNA-sequencing profiles RNA from intact cells, while single-nucleus RNA-sequencing profiles RNA from isolated nuclei. scRNA-seq is often used for fresh cell suspensions, while snRNA-seq is commonly used for frozen, fragile, or hard-to-dissociate tissues.
What should a nucleus isolation protocol include?
A nucleus isolation protocol should include cold handling, tissue disruption, selective cell membrane lysis, filtration, washing, debris control, nuclei counting, and final concentration adjustment for the sequencing platform.
Conclusion:
Single-nucleus RNA-sequencing is a practical approach for profiling long-term frozen brain tumor tissues, including pediatric glioma samples and other challenging tumor specimens. A simplified workflow focuses on cold handling, gentle tissue disruption, selective lysis, filtration, washing, debris cleanup, nuclei counting, and platform-ready preparation. Compared with single-cell RNA-sequencing, snRNA-seq is especially useful when fresh intact cells are unavailable or difficult to recover. By choosing nucleus isolation and sample preparation solutions that match the tissue, storage conditions, and downstream application, researchers can build a more reliable workflow for studying tumor heterogeneity and molecular biology at single-nucleus resolution.